We have a new manuscript from the lab in Cell Reports, Distinctive Synaptic Structural Motifs Link Excitatory Retinal Interneurons To Diverse Postsynaptic Partner Types. This manuscript is in collaboration with the first author, Wan-Qing Yu @wanqing_yu, then co-authors Rachael Swanstrom, Crystal L. Sigulinsky @CSigulinsky, Richard M. Ahlquist, along with Sharm Knecht, myself Bryan W. Jones @BWJones. David M. Berson, and Rachel O. Wong. The PDF is here.
Abstract:
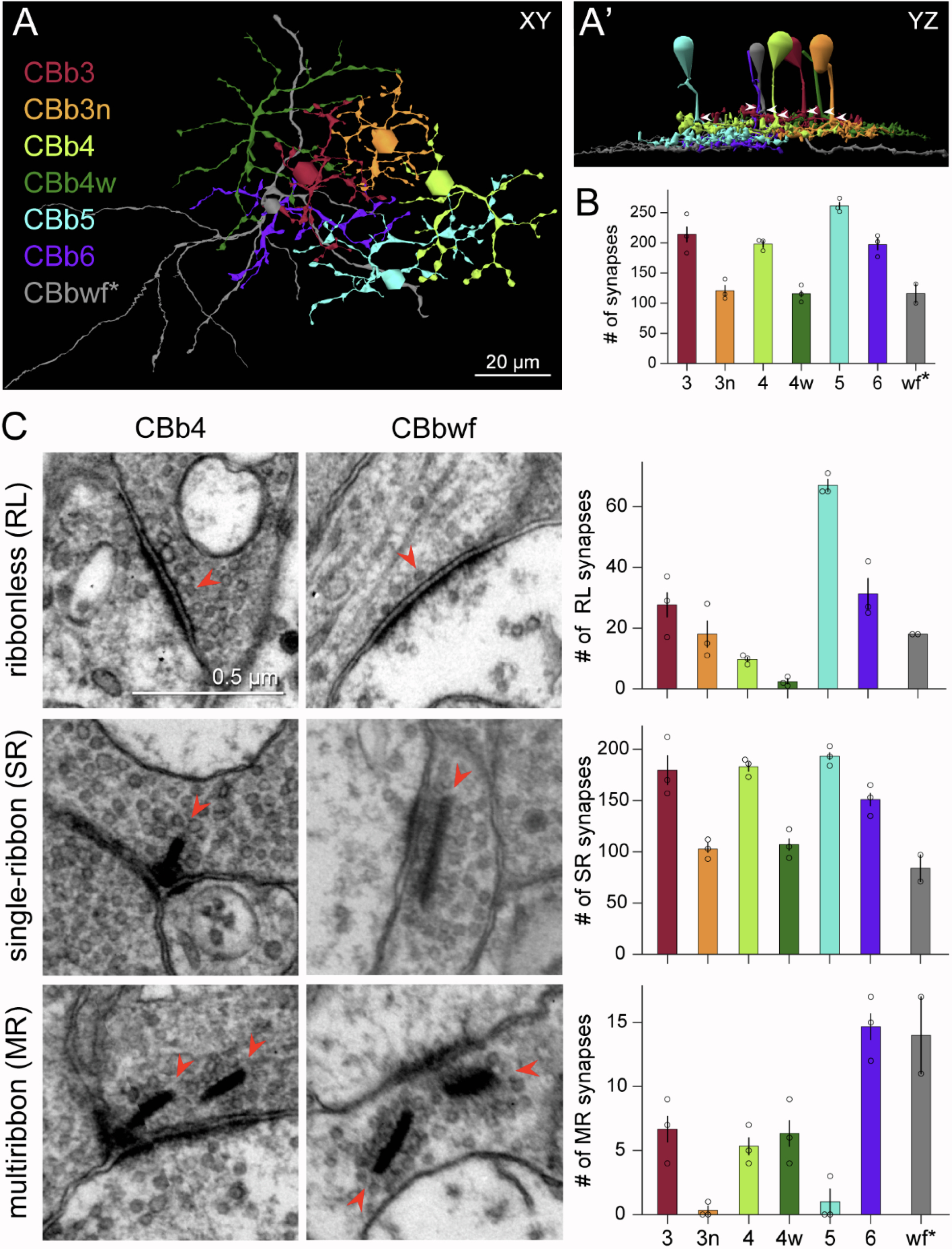
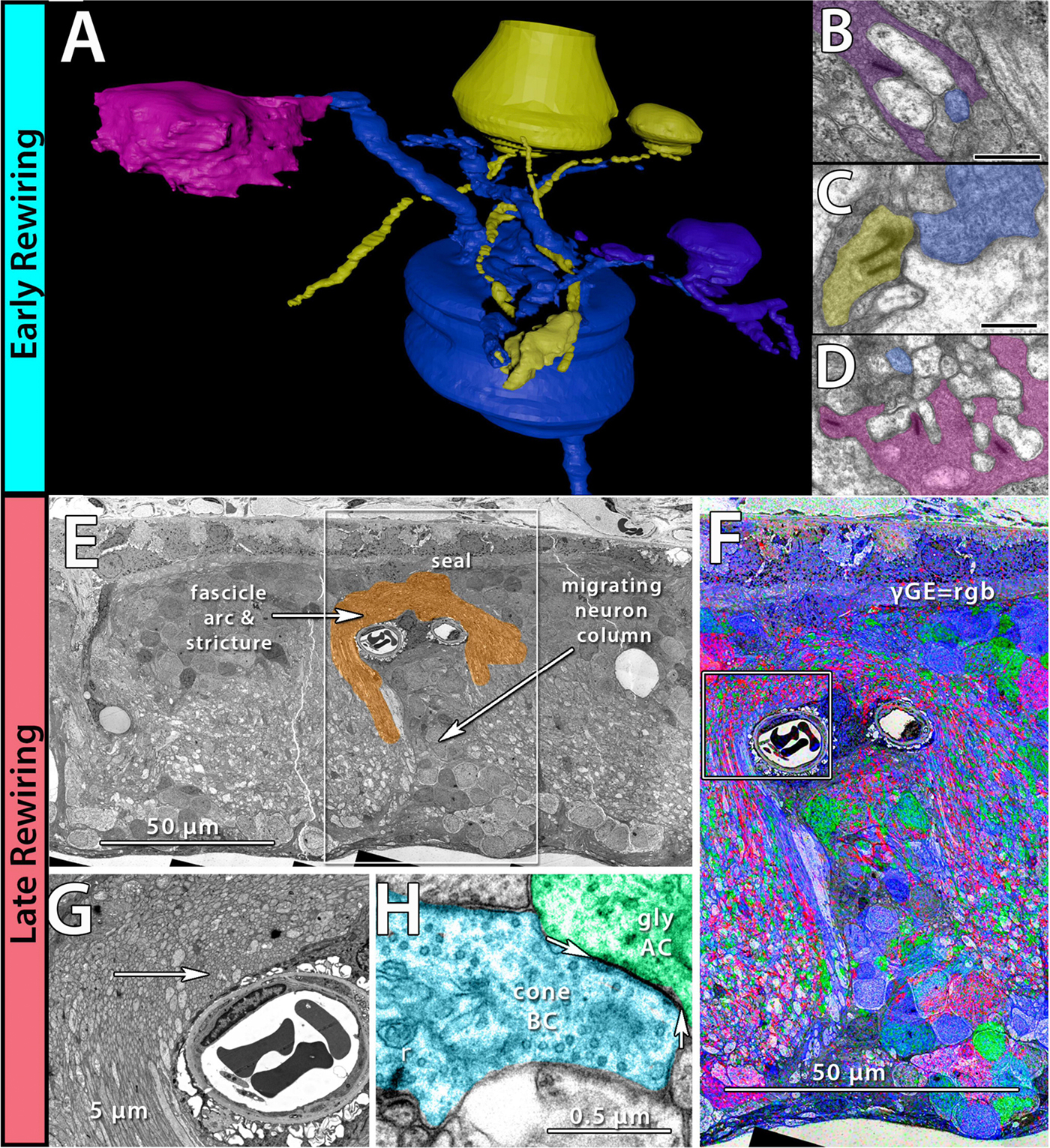
Neurons make converging and diverging synaptic connections with distinct partner types. Whether synapses involving separate partners demonstrate similar or distinct structural motifs is not yet well understood. We thus used serial electron microscopy in mouse retina to map output synapses of cone bipolar cells (CBCs) and compare their structural arrangements across bipolar types and postsynaptic partners. Three presynaptic configurations emerge—single-ribbon, ribbonless, and multiribbon synapses. Each CBC type exploits these arrangements in a unique combination, a feature also found among rabbit ON CBCs. Though most synapses are dyads, monads and triads are also seen. Altogether, mouse CBCs exhibit at least six motifs, and each CBC type uses these in a stereotypic pattern. Moreover, synapses between CBCs and particular partner types appear biased toward certain motifs. Our observations reveal synaptic strategies that diversify the output within and across CBC types, potentially shaping the distinct functions of retinal microcircuits.